General
WildTrax is an online platform for managing, storing, processing, and sharing biological data collected by environmental sensors. WildTrax provides tools for managing large data sets, and also creates opportunities to address broad-scale questions using novel approaches. Supported environmental sensors currently include autonomous recording units (ARUs) and wildlife cameras, while a third sensor, point counts, provides access to manage avian count data collected by humans.
WildTrax is available for organizations using cameras and/or ARUs—simply create an online account to begin. Contact info@wildtrax.ca or see The Guide on how to get started.
WildTrax offers several benefits for users of environmental sensors, including:
- Improvement in processing speed
- More accurate, and higher quality data using the species verification
- Unlimited, online access to your data
- Seamless and flexible data sharing options with teams and collaborators
- Standardized approaches to data collection across networks of organizations and individuals
- Opportunities to discover data in your region of interest, coordinate with other groups, and address broad-scale ecological questions
- Centralized repository for long term data archiving
Sensors can be deployed in many ways, depending on the monitoring objective. Examples of methods and protocols are found here.
The set-up methods used by the ABMI are summarized below:
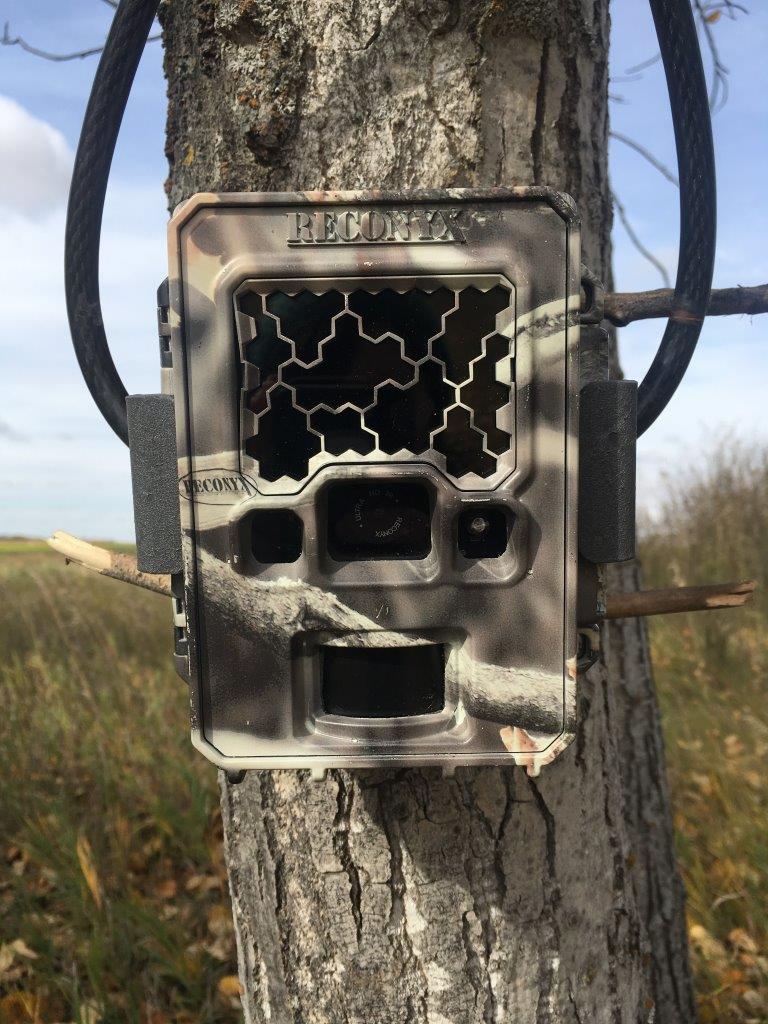

For cameras, choose a view that is not blocked by vegetation or other impediments for at least 10 m (try to anticipate vegetation growth). Set the camera (lens) height at 1 m and then focus the camera view on the reference stake at 80 cm above the ground. Your target detection zone should be approximately 3–5 m from the camera. Face the camera to the north (ideally) or south if possible to avoid visibility issues from direct sunlight.
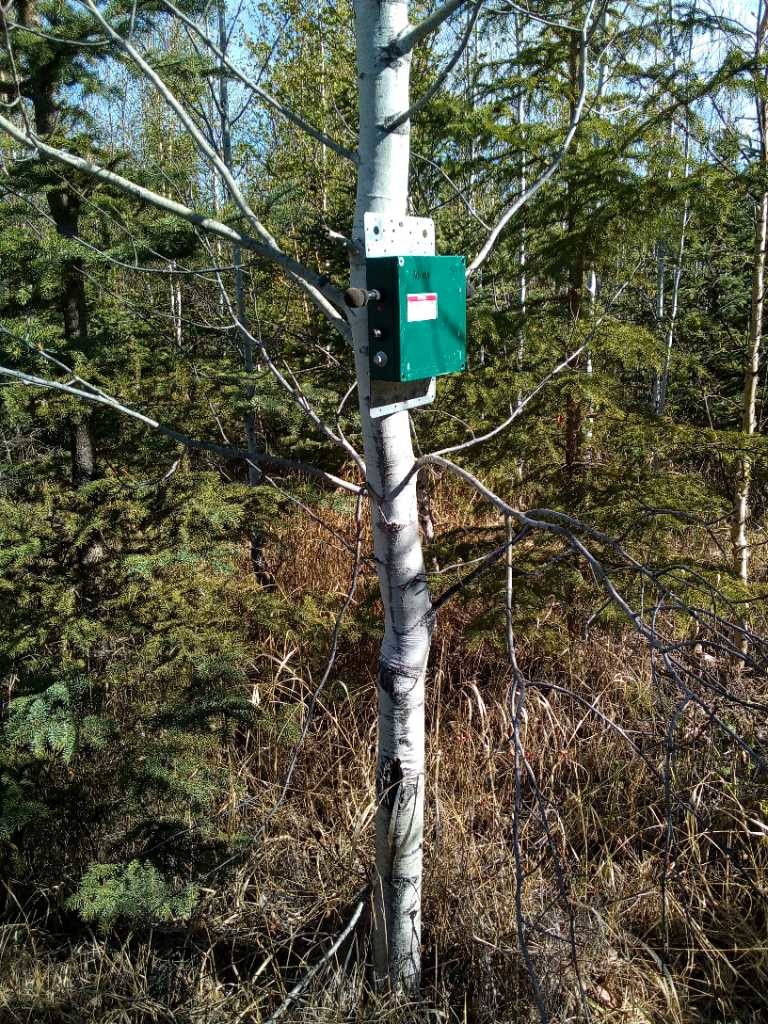

The autonomous recording unit (ARU) should be at a height of 1.5 m above ground, facing north with the microphones unobstructed by leaves, branches, or (if applicable) the trunk of the tree to which it’s affixed. Choose a sturdy tree or support, such as a stake, so that the unit won’t topple over in high winds or if disturbed by a large mammal.
Creating an account, setting up an organization, and processing and publishing projects are all free.
Exceptions may occur if an organization attempts to upload tens of millions of images or recordings per year, in which case there may be some required cost recovery for the storage fees on AWS, where the media are stored.
Your upload and download speed are limiting factors when you interface with WildTrax. The platform attempts to resolve these issues by providing asynchronous uploading and caching large data sets, but be mindful of the volume of data you’re uploading and downloading and your internet connectivity before proceeding.
Remote Cameras
Currently, WildTrax has not encountered a make and model of camera that it does not support. However, it cannot populate data from fields that do not exist in the image Exif data. Metadata fields are stored in the image metadata e.g., “Sequence” or “TriggerMode”; however, the fields stored as image metadata or “Exif” data vary by camera make and model. If it seems like you are having issues due to the make or model of your camera, please contact support@wildtrax.ca. You can also refer to Phil Harvey’s ExifTool Tag Names to determine which tags are available for your camera make and model.
If you are purchasing remote cameras for the first time, Reconyx cameras (e.g., HF2, HF2X, PC800, PC900, and HC600 models) are greater for first-time users as they are user-friendly and intuitive.
High-quality SandDisk SD cards or Kingstone Class 4 and 10 SD cards are frequently used. We would not recommend anything below a Class 4 write speed.
For more information on camera brands, please consult the Remote Camera Survey Guidelines.
WildTrax integrates the following:
(1) Microsoft’s Megadetector v5, that automatically tags images of vehicles, animals, humans or NONE.
(2) Microsoft’s MegaClassifer v0.1.
(3) A “Staff/setup” tagger designed to filter out images of humans at the camera’s deployment and retrieval. When enabled in your project’s settings, these tools will automatically filter and tag your images.
This results in less time spent sifting through false fires and more time spent focusing on the species you want to tag.
Cameras can sometimes capture images that do not contain wildlife—‘false fires’—due to movement in vegetation or changes in sunlight. These false fires can increase processing cost and time. To aid in processing these images, WildTrax contains a model to automatically identify false fires, allowing them to be removed before further processing. The model uses training data from 1,325 camera deployments as well as a trained network, CaffeNet, specifically modified for WildTrax. This tool results in less human time spent sifting through images of vegetation movement. The model was validated with an additional 121 camera deployments with 79,451 false-fire images. The model identified 34,456 (43.6%) of false fires with a 0.2% error (false positive) rate. That is, more than 40% of false fires can be reliably (0.2% error) removed before processing. Depending on the camera unit used, image quality and habitat type results may vary.
Images cannot be deleted individually in WildTrax (including those of humans); instead, WildTrax allows you to filter images of humans using the results from Megadetector (if enabled in project settings) as well as select options to opt-in for human blurring (can be enabled in organization settings).
ARUs
Wildlife bioacoustics is the study of animals using the vocalizations that they produce. Sounds are identified to the species or even individual level using unique patterns known as spectral signatures. These data are used to answer research and monitoring questions about individual species or groups of species.
The Bioacoustic Unit is a collaboration between the Bayne Lab at the University of Alberta and the Alberta Biodiversity Monitoring Institute. Our research group develops tools, protocols, and recommendations for acoustic monitoring programs across the country.
To learn more about the Bioacoustic Unit, please click here.
The BU uses robust environmental sensors, called Autonomous Recording Units (ARUs)—essentially sophisticated battery-operated microphones—to record sounds produced by vocalizing animals. There are recommended settings that can be used to optimize recordings of birds, mammals, and other taxa.
The Bioacoustic Unit uses Song Meter Autonomous Recording Units made by Wildlife Acoustics. Most of our Song Meters are the SM2+ and SM4 models. Other less frequently used models include the SM3, the SM2 with GPS, and the SM2+BAT. The GPS-enabled units permit more precise localization of animals in space. For memory cards, we often use high-quality SanDisk SD cards. We also occasionally use the Kingstone Class 4 and 10 SD cards. Wildtrax can take data from any type of digital sound recorder.
Cumulatively, more species are observed by going to new stations within a study area than by listening to more recordings of the same locations; however, the difference is not that large. If sufficient funding exists to go to more locations, that will provide a better estimate of total species. However, when restrained by field costs, leaving ARUs in the same location and repeatedly sub-sampling is recommended, particularly if you are interested in multiple taxa (e.g., owls and songbirds).
For songbirds, leaving an ARU out for several days will yield higher occupancy rates and probability of detection than repeatedly sampling in a single day. The additional benefit of leaving an ARU out for a month is relatively small for songbirds. However, there is evidence that more species will be detected with more sampling effort and owls, amphibians, and mammals have very different calling behaviours from songbirds.
Minimum sampling effort recommended by the BU in order to maximize detection for most acoustic species is 3–7 days. Each sampling event is recommended to be at least 3 minutes long, either at dawn or dusk and at least one day apart.
The question here is whether you could achieve the same results by listening to the same total number of recordings from a single day vs. a week vs. a month. Sampling for approximately a week results in higher estimates of species richness at a station compared to sampling for a day. In our tests, there was no significant difference between leaving an ARU out for a week vs. a month but that was only for songbirds.
This is entirely dependent on the frequency with which a species sings. The Bioacoustic Unit and the Boreal Avian Modelling project have estimates for all species, however, so you can assess the effort required to ensure you detect a species if it is present.
Calling rate has the greatest effect on detection rate, explaining 49% of the variance in detection rate. Calling rate coupled with the abundance of a species, time period, and a species’ log body weight explained 69% of the variance in detection rate. When the abundance of a species is high, there is higher detectability. Species that call at night have lower detection rates than those that call during the day. Also, larger species generally have lower calling rates. In general, species that are less abundant, have a large body weight, and vocalize infrequently and/or more often during the night have a lower detection rate and will require more extensive sampling.
There are consistent benefits to repeatedly sampling at the same station when estimating trends for a species as you are more certain if the species is present or absent. However, the statistical power of trends is driven by the number of stations and the number of years observed.
Within the first minute of a 10-minute point count, 49.8% of all vocalizing species are detected. Within the first five minutes, 79.2% of all vocalizing species are detected. However, if you have the choice between 10 1-minute samples taken at different times of day or year and 1, 10-minute period you will detect far more species using 10 1-minute segments.
Using more point counts with shorter duration detected a larger proportion of all species compared to fewer, longer duration point counts.
If you sample only a few points from the total number of available recordings, there is strong evidence that afternoon sampling can be avoided altogether if you are relying on listening.
Recognizers can be used when you are targeting a specific species, and a manual scanning spectrogram can be very effective in processing data when vocalizations are visually distinctive and recognizable. In short, the training data is used to create a template (“recognizer”) and is then matched to a recording segment from the test data. More information can be found here.
BirdNET is a multi-species bird classifier developed by Cornell University. WildTrax utilizes the BirdNET API to allow users to obtain results from the classifier for their projects.
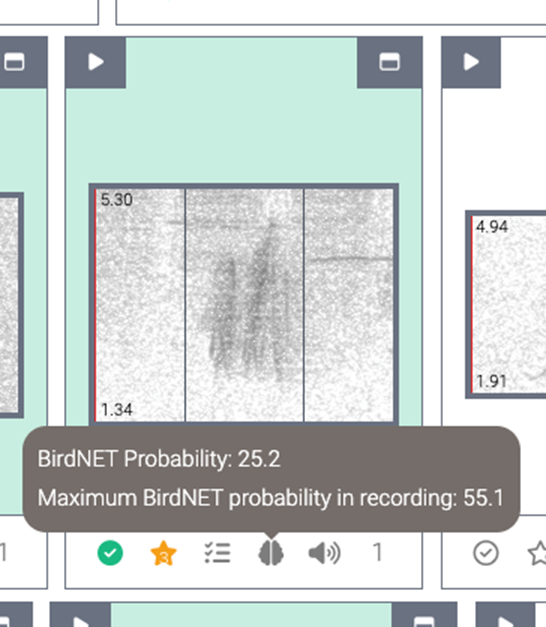
When using BirdNET as a part of species verification, you can identify whether or not the tag achieves a high BirdNET confidence level by hovering over the brain icon. This is usually useful when the time of first detection of the tag is not high quality; however, if the individual is occupying the space around the recorder with high enough signal amplitude and unobstructed calls, BirdNET’s confidence will be more valuable.
You can also use BirdNET as a guide to automatically return the species it thinks it found. BirdNET provides values in 3-second windows for each recording in a project. You can find BirdNET’s output in the _birdnet_report.csv in Data Downloads.
Evaluating the performance of BirdNET on a data set is also possible. In a binary classification task (BirdNET = predicted, human = observed), you can distinguish false positives (incorrect detections) and false negatives (missed detections) and subsequently create performance metrics. This is useful for questions like species presence, where the highest confidence value of BirdNET across many recordings, can yield a positive result, with minimal effort.
Data Management and Privacy
You must first create an Organization to house your data and metadata. The WildTrax team will review your request and confirm your identity after which point you will have the ability to create projects under your newly created Organization and upload media into those projects.
There can only be one Organization for a Project. You must choose who owns that data. This must also be the organization who collects and uploads the data. The owner of the data can then add the collaborating users to their project or organization pending on the level of collaboration.
In your Organization settings, change all locations to the specific buffer you are comfortable with. For each project you wish to share choose “Published – Map + Report” as non-authorized users will not be able to see the media or the un-buffered location this way.
WildTrax uses various permission and privacy levels throughout the system to ensure your data is secure to the level you choose
WildTrax has two basic membership levels : Admin and Read-Only. See below for more details on what each membership level can do
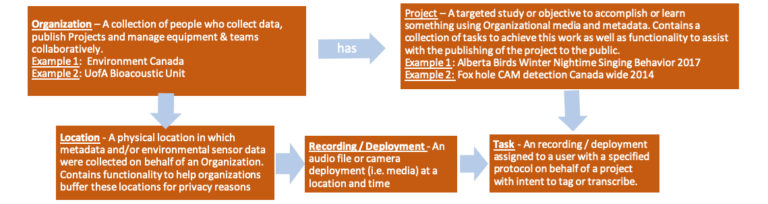
Organizations are the framework of your data on WildTrax. They contain your locations, visits, equipment and summaries of your acoustic recordings and camera data upload to your Projects.
An organization usually contains a collection of people who collect data, publish projects and manage equipment and teams. Administrators can read and write un-buffered locations on all organizational locations, add users to Organizations and Projects. read and write location, visit and equipment metadata and inherit administrator privileges on all organization projects. Read-Only members have read un-buffered on all organizational locations, can read location visit metadata and equipment and can read all organization projects.
Projects are a targeted study or objective to accomplish or learn something using environmental sensor data. Projects are populated using locations, visit and contain a collection of tasks to achieve this work. Project administrators manage projects, assign tasks and users, and publish projects. Read-Only members can view and download data depending on their level of access via the Project Privacy Levels: Active; Test Only – Hidden; Published – Private; Published – Map + Report Only; Published – Public.
Choose Published – Private if you don’t want to receive access requests or Active if you are still working on completing your project.
Locations are a physical location associated with the deployment of an environmental sensor. Locations contain subsequent visit and equipment metadata that were collected on behalf of the organization. Locations can be either hidden, buffered, true location + buffer and true location. This allows for flexibility with specific locations requiring a higher level of privacy.
Tasks are recordings or deployments assigned to a specific user and tagging protocol. Read-Only members can gain privileges to write at the task level in order to perform tagging. Organization or project administrators can also change the tags in the tasks, which is subsequently tracked in the audit table
Recordings (ARUs) / deployments (cameras) are audio files or a series of images corresponding to an ARU / camera deployment.
For each project make sure you choose Published – Private when it is complete. To give someone access to a project, add the user into the Project as a member (this will give access to just the current project + buffered access) or add the user as a member into the Organization (this will give access to all projects + un-buffered access). You can also remove members from the project or Organization or reduce their access at any time.
Active
Test Only
Published – Private
Published – Map + Report Only
Published – Public
Buffer your locations in organization settings. This tells WildTrax this location is already buffered and no one will know the true location because WildTrax was never given that information.
You must add yourself as a project member to see the projects in the default view. You can hit the “View Only My projects” toggle in the top right to see all projects you are eligible to see. WildTrax does this to try to help simplify the amount a user sees at once as it would be overwhelming and most of the time people are only interested in their own projects.
Organization investigators are the ones who receive all access requests. Requests will only go to the administrators of the project/organization if no investigators have been assigned.
Make sure your Organization has an investigator set to handle all of these requests. You can do this in organization settings.
All data uploaded to WildTrax is by default private and only viewable by the project members depending on their level of membership (Admin or Read-Only). Project and organization administrators maintain ownership and privacy rights over uploaded data, regardless of whether the data is private or publicly available. When you upload data to WildTrax, you have the option of releasing your data publicly or not.
